Media
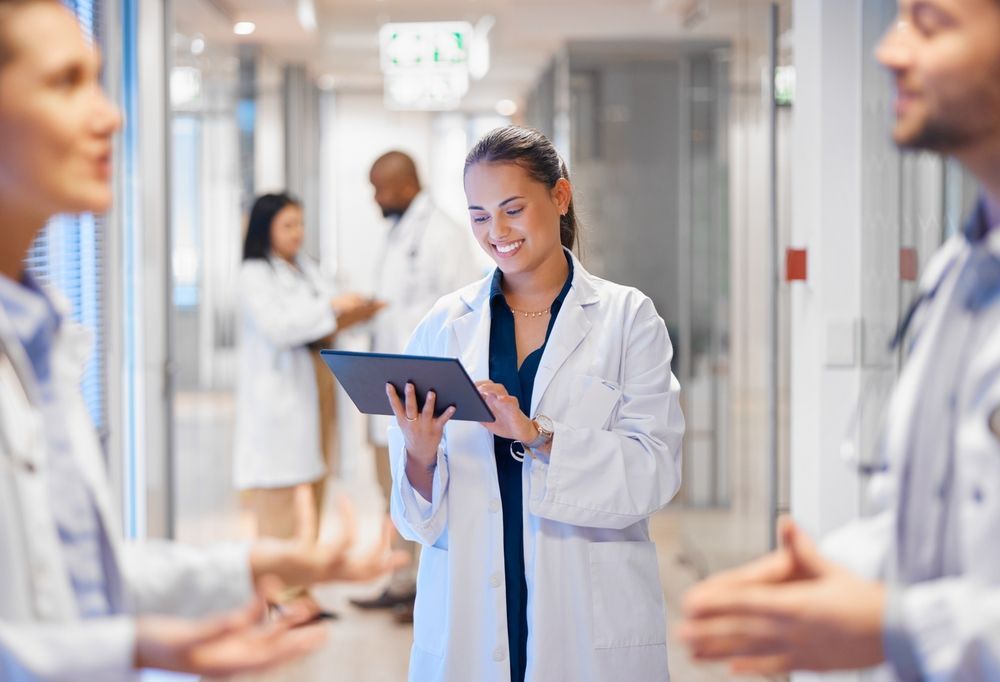
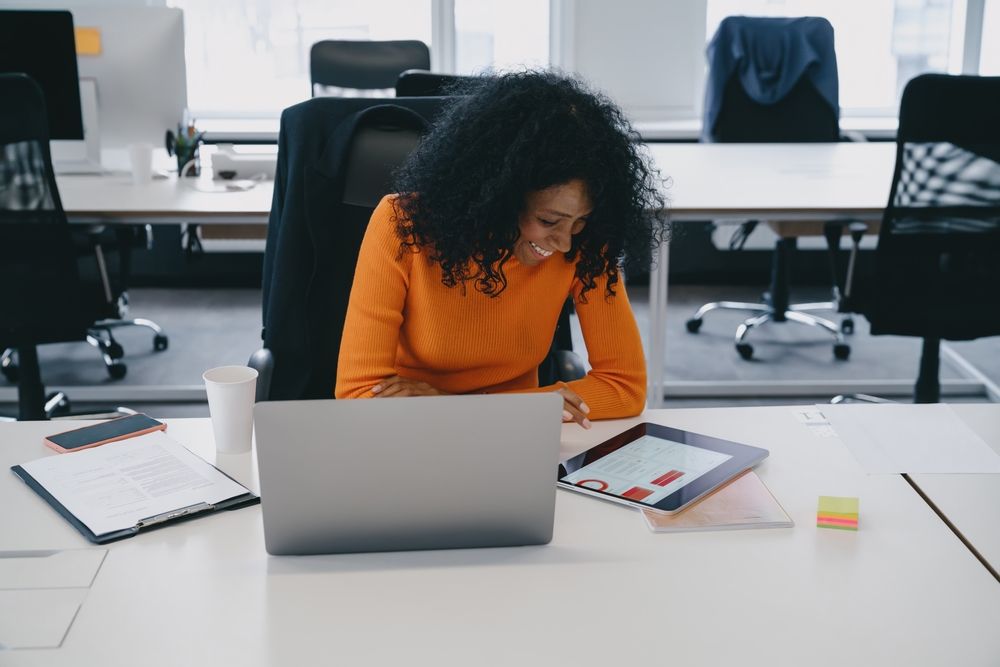
Healthcare Insights from Professionals
We constantly leverage our extensive industry knowledge and expert relationships to reveal the latest trends across MedTech markets in order to bring you actionable news and insights.
Latest Articles
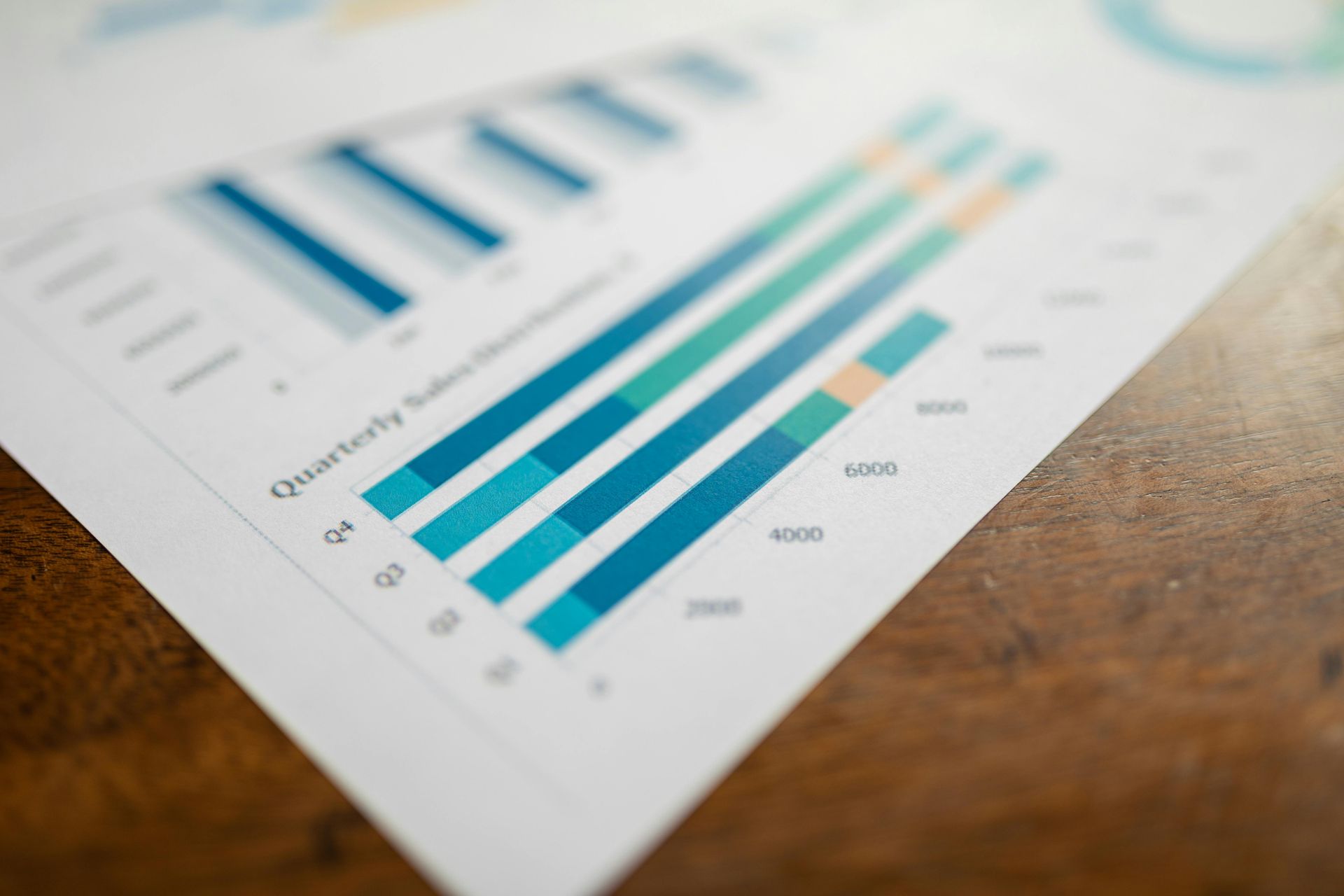

Explore the Q4 2025 IVD Performance Scorecard to see how major acquisitions and structural shifts are redefining the global diagnostics industry. Analyze how 11 leading companies are navigating China’s procurement reforms, the shift toward specialty assays, and the high-stakes race in rapid antimicrobial susceptibility

Contact Us
We would love to hear from you. Send a message via our contact page and we will get back to you as soon as we can.